 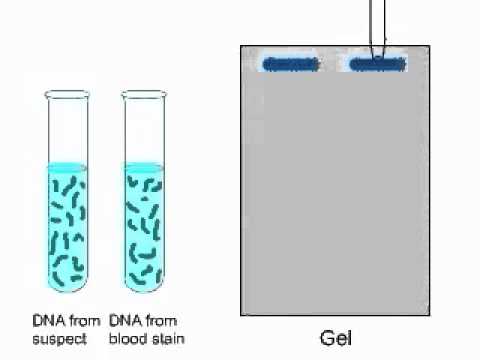
Each of these enzymes cleaves DNA between different, and specific, sequences of nucleotides. 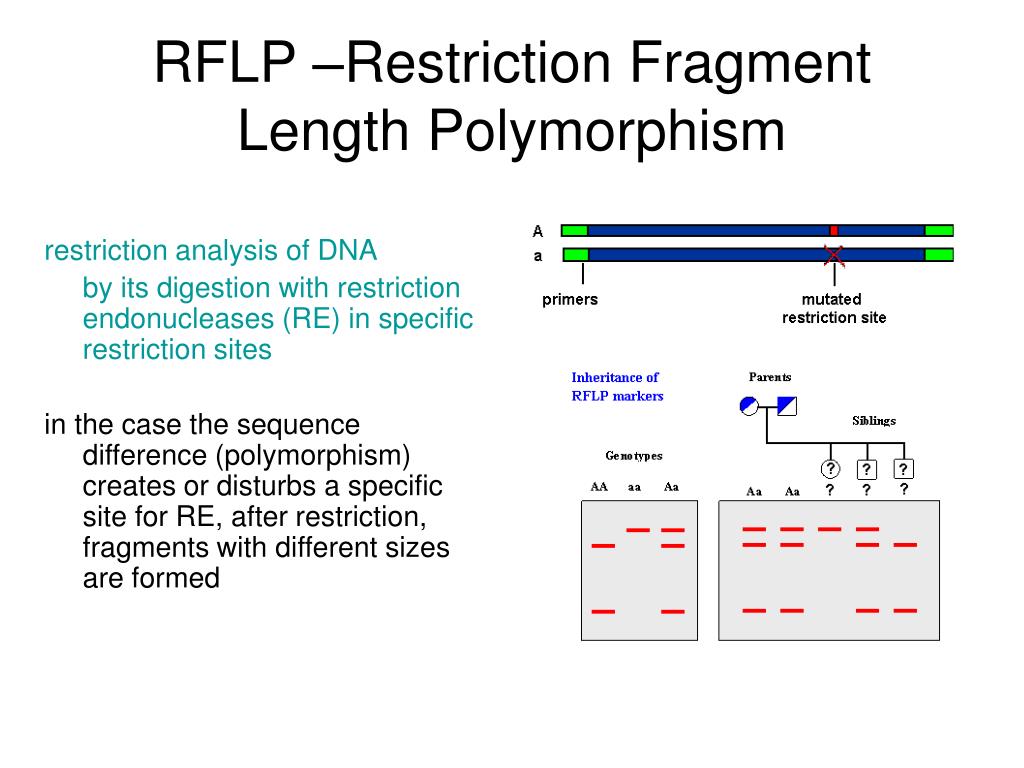
Over 90 different restriction enzymes have been isolated from different species of bacteria. Bacteria naturally produce restriction enzymes and they use them to cleave the DNA from foreign organisms. For example, the restriction enzyme HaeIII recognizes the sequence GCGC and it cleaves the bond between middle cytosine and guanine. This is simply the replacement of a single nucleotide by a different one.Ī special type of protein called a restriction enzyme, or a restriction endonuclease, can recognize specific sequences of nucleotides on DNA and then cleave the DNA at these locations. Two families of these repeats are found quite often in DNA: variable number of tandem repeats (VNTRs) and short tandem repeats (STR). In vertebrates (animals with a backbone), there are regions of DNA that contain many repetitions of the same sequence. Deletions are regions where nucleotides have been removed. Insertions are regions of DNA where nucleotides have been added to a sequence. There are a variety of types of mutations in DNA. Differences in individuals result from small variations, called mutations, in the sequence of DNA. The sequence of these nucleotides is extremely important, as it determines the structure of all of the molecules in an individual. The four nucleotides are adenine (A), guanine (G), cytosine (C), and thymine (T). The DNA molecule is made up of a sequence of four smaller molecules called nucleotides. RFLP is used as part of DNA fingerprinting, to detect genetic diseases and to determine genetic relationships between species. Electrophoresis is then used to separate the strands according to their length. Mutations within the DNA result in strands of different lengths. Special enzymes that cleave the DNA in specific locations are used to digest strands of DNA. RFLP, or restriction fragment length polymorphism, is a molecular biological technique used to compare DNA from two samples. The method appears to be viable for rapid analysis of mycorrhizal communities using pooled root tips collected from soil cores.RFLP (Restriction Fragment Length Polymorphism) We found good correspondence between the community distribution patterns, determined by T-RFLP, and the individual morphotype T-RFLP patterns normalized by their QPCR values. DNA extracted from morphotype classes from the same core were pooled, amplified as above, and used to assess abundance of fungal species types within the mixed community and results compared to traditional morphotyping techniques. Restricted labeled amplicon was used to generate terminal restriction fragment lengths on an ABI 310 sequencer to determine molecular identity of colonizing fungi. Root tips separated from soil cores were morphotyped according to standard procedures, and DNA was extracted from separate morphotype classes and amplified with a novel set of primers labeled with the fluorescent dyes 6FAM and HEX. We describe here results of a study conducted to assess the efficacy of a T-RFLP approach for determination of ectomycorrhizal communities colonizing the roots of Loblolly pine (Pinus taeda L.) in North Carolina, U.S.A. T-RFLP methodology targeting rRNA genes has effectively been used to discriminate between microbial communities but to date has not been used specifically for the analysis of ectomycorrhizal communities colonizing plant roots. Consequently, miscalculation of indicators of community structure (richness, diversity indices, evenness) can occur and the ability to address ecological/ecosystem function questions is curtailed by the high sample time commitment. Depending on the rigor of the classification protocol, this technique can incorrectly assign dissimilar genetic entities into the same morphotype class and is very labor intensive. Studies of ectomycorrhizal community structure have used a variety of analytical regimens including sole or partial reliance on gross morphological characterization of colonized root tips. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
